Polycomb repressive complex 2 (PRC2) is an enzyme complex with wide reach, found in multicellular organisms from plants and flies to humans, and which is essential to the development and livelihood of the whole organism. It is critical to maintaining cell identity by keeping thousands of genes inactive.
Mutations in genes coding for PRC2 subunits or their overexpression are associated with most types of cancer, making the enzyme complex of considerable interest to cancer clinicians and researchers.
In a major development for the field, an EMBL Australia Group Leader has revealed exactly what happens at the site where PRC2 interacts with RNA, the immediate product of genes. For Associate Professor Chen Davidovich, who has studied the way PRC2 is regulated by RNA for close to a decade, this was like “closure”.
The study, published in Nature Structural and Molecular Biology, was conducted in collaboration with the laboratory of Associate Professor Roberto Bonasio (University of Pennsylvania) and Dr Ralf Schittenhelm who heads the Monash Biomedical Proteomics Facility.
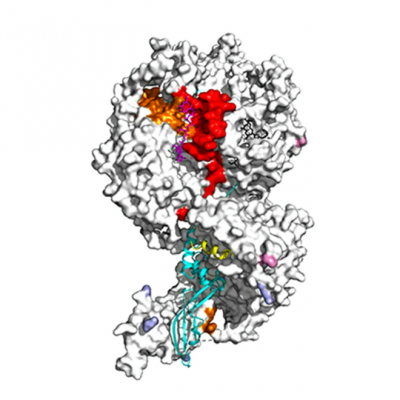
It has been known for more than a decade that PRC2 binds tightly to RNA and it was later shown that RNA enervates its enzymatic activity. Yet it was unknown exactly how this happened and, importantly, under what circumstances the inhibitory activity of RNA is relieved. The researchers used mass spectrometry to tease out the processes at play, then backed up their discoveries using other methods. Perhaps surprisingly, they found that the factors that inhibit PRC2’s enzymatic activity also occur at the same site on the complex (its regulatory centre) as those that stimulate it.
“Overall this study helps to explain how the positive and negative effectors of PRC2 regulate enzyme activity – I think this is very significant in the context of the regulation of PRC2,” Associate Professor Davidovich said, who is also a member of Monash Biomedicine Discovery Institute (BDI) and the ARC Centre of Excellence in Advanced Molecular Imaging.
“I’m really excited about it; I’ve really wanted to know where the site was for many years,” he said.
It’s a question many different research groups have tried to answer.
“PRC2 doesn’t have a classical RNA binding domain, which has meant that so far you had to guess where RNA might bind.”
“What we think – and this is only a model – is that RNA is a sort of safety switch to prevent PRC2 from working in genes where it shouldn’t.”
Associate Professor Davidovich credits the “outstanding” Monash Biomedical Proteomics Facility with providing the tools that were key to the discoveries.
Postdoctoral researcher Dr Qi Zhang and PhD student Nick McKenzie were first authors on the paper, while Associate Professor Davidovich was the senior author.
Read the full paper in Nature Structural & Molecular Biology titled RNA exploits an exposed regulatory site to inhibit the enzymatic activity of PRC2. This article was published on the Monash BDI website and republished here.